Cationic Liposome (Genesome®) containing DDAB (Plain and Fluorescent)
Description
Cationic liposomes are traditionally used for the delivery of genetic materials such as various types of DNA (pDNA, cDNA, CpG DNA, oligonucleotide, antisense oligonucleotide, etc.), various types of RNA such as (siRNA, mRNA, etc.) and nucleic acid mimics (NAMs). The encapsulation of DNA into the conventional neutral charged PC based liposomes can be a technical problem mainly due to the plasmid size. Due to this problem in late 80s, the liposomes composed of cationic lipids and PE have been developed. The idea was to neutralize the negative charge of pDNA with positive charge of cationic lipids in order to capture more plasmid efficiently mainly due to electrostatic interaction and deliver them into the cells. Generally, the procedure is simply based on mixing the cationic liposomes with DNA or RNA and adding them to the cells. This results in the formulations of aggregates.
In order to design a proper cationic lipid for gene delivery, two approaches have been used for the cationic lipid synthesis: 1) cholesterol-based design such as DC-Cholesterol and GL-67 lipids, and 2) non-cholesterol-based designs such as DOTAB, DDAB and DOTMA. To successfully transfer the gene in vitro using liposomes, some consideration should be taken into account: i) the ability of binding and packing DNA/RNA in liposomes; ii) the interaction of the packaged DNA/RNA to the cell surface; iii) the efficiency of the internalization of DNA/RNA; iv) the intracellular DNA-release from endosomes in case of endocytosis involvement; v) the transgenic expression level in cell nuclei. pH-sensitive liposomes have been designed based on their tendency to release their content in the acidic condition. The primary concept was based on viruses that fuse with the endosomal membrane by means of a protein at pH 5-6, delivering their genetic material to the cytosol before reaching the lysosomes. Typically, a pH-sensitive liposome consists of dioleoylphosphatidylethanolamine (DOPE). Since phosphatidylethanolamine (PE) changes in acidic conditions, it is believed to act as a membrane fusion promoter. The effectiveness of the interaction between liposomes and cells is highly dependent on the liposome compositions. Liposomes are captured by various endocytosic processes, and the efficiency depends on the cell type and liposome size. Liposomes of various sizes and charges can attach to the macrophages and neutrophils through active phagocytosis. After attachment of the liposome to the cell surface, the internalization into the endosomes occurs due to a more acidic pH (6.50) at early endosomes. The liposomes are transferred to the last endosome with more acidic pH (5.5-6.0) by maturation or vesicular fusion, which takes 10-15 min. Twenty minutes (or more) after uptake, the contents are delivered to the lysosome with pH 5.0 or less. Lysosomes are the main degrading and last endocytotic section in the endocytotic pathway, in where pH-insensitive liposomes are accumulated and degraded. However, after penetration of pH-sensitive liposomes into cells, the accumulation and degradation do not occur.
Formulation Information
Cationic Liposome (Genesome®) containing DDAB (Plain and Fluorescent)
For more information on the lipid composition of the liposomes mentioned above click here.
| Buffer and Liposome Size | Specification |
|---|---|
| Buffer | Deionized RNAse-free Water |
| pH | 7 |
| Liposome Size | 100 nm |
Fluorescent Dyes for Fluorescent Cationic Liposomes
| Fluorescent Dye | Excitation/Emission (nm) | Molecular Structure |
|---|---|---|
| 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-(7-nitro-2-1,3-benzoxadiazol-4-yl) (ammonium salt) | 460/535 | 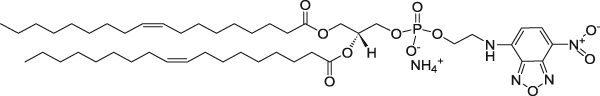 |
| 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine-N-(lissamine rhodamine B sulfonyl) (ammonium salt) | 560/583 | 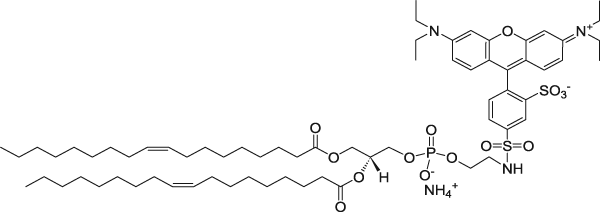 |
Nitrogen/Phosphate (N/P) Ratio
The ratio of positive-to-negative charge was calculated based on the cationic amine nitrogen (in the cationic lipids) and DNA phosphate group concentration. DNA and RNA are a polymer of nucleotides. They join themselves through phosphodiester bonds (a specific kind of covalent bond) that can grow up to millions of nucleotides. DNA and RNA are negatively charged due to the existence of phosphate group that makes up every nucleotide (pentose + nitrogenous base + phosphate). When forming part of the phosphodiester bond, they retain 1 of 2 negative charges. The other negative charge is lost to form the other ester bond to a new pentose. That is the reason the bond is called “phospho-di-ester”. As an example, cationic lipid DDAB has the following structure.
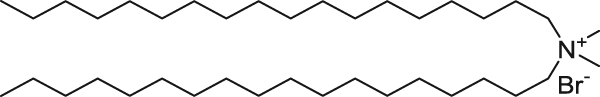
DDAB has one nitrogen and therefore, there is one positive charge per MOLECULE. 1 nmol of DDAB contribute to 1 nmol of positive charge. Some cationic lipids contain more than one nitrogen, but not all nitrogen atoms carry a positive charge. For other lipids, look at their molecular structure to find the number of nitrogen atoms in the molecule.
siRNA Liposomes
RNA, unlike DNA, is a single stranded molecule. However, siRNA is double stranded. As an example, in an siRNA molecule that is 21 bp long, there are 42 negative charges since there are 2 nucleotides in a base pair and each nucleotide carries one negative charge.
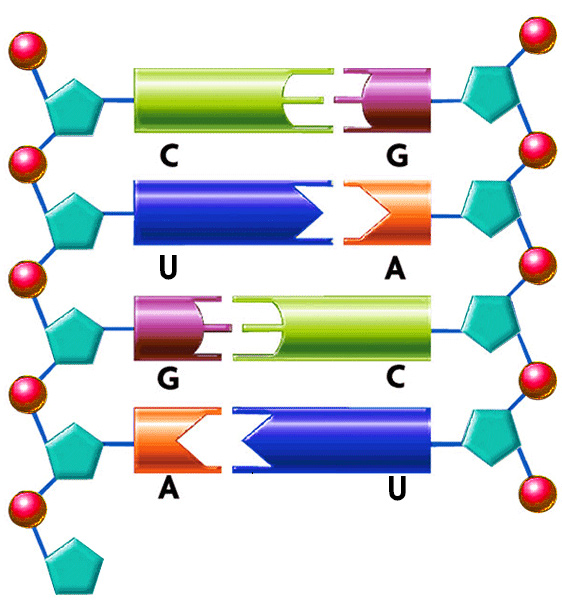
DNA Charge Calculation
1 µg of DNA contributes to 3.1 nmol of negatively charged phosphate.
Cationic Lipid–Protamine–DNA (LPD) Complexes
Protamine is a highly positive charged polycationic peptide with a molecular weight of 5.1 kDa that acts as a DNA condensation reagent. Protamine is known to be the major component in the sperm nucleus for condensing DNA. All protamines contain arginine and are strongly basic (isoelectric point = 11-12).
Lipopolyplexes composed of lipid-Protamine-DNA (LPD) are generated by the association of protamine pre-compacted DNA with cationic liposomes. This is in contrary to cationic liposome-DNA lipoplexes that are formed by the direct electrostatic interaction between cationic lipid and DNA. Lipid-Protamine-DNAs have a net positive surface charge, believed to be essential for their binding to anionic cell surface proteoglycans. This electrostatic interaction at the cell surface triggers lipid-DNA complex endocytosis into the cell.
The order of adding protamine to DNA and cationic liposomes happens to matter a lot. One protocol involves precomplexation of plasmid DNA with protamine sulfate followed by the addition of cationic liposomes. There are two disadvantages with this protocol. One of them is that large amounts of excess cationic lipids are required to achieve a maximal level of gene expression. Typically, a cationic lipid/DNA molar ratio of greater than 35 is necessary for an efficient in vivo transfection. The other problem is the difficulty in preparing a concentrated sample. This is due to the overwhelming interaction of protamine sulfate with plasmid DNA which makes the preparation of protamine/DNA complexes difficult at high concentrations, especially when a high protamine/DNA ratio (w/w) is required. To solve these problems, a second protocol has been developed, in which protamine sulfate is mixed with cationic liposomes followed by the addition of plasmid DNA. Due to the competition between cationic liposomes and protamine for the interaction with plasmid DNA, the strong interaction between protamine and DNA is greatly reduced. Under optimal conditions, this will result in formation of LPD with a mean diameter less than 150 nm.
In order to calculate the positive to negative ratio, you need to consider:
- 1 mol of protamine sulfate contributes to 21 mol of positive charge (MW of protamine sulfate is about 5.1 kDa).
- 1 nmol of a monocationic lipid contributes to 1 nmol of positive charge.
- 1 µg of DNA contributes to 3.1 nmol of negatively charged phosphate.
Post-insertion PEGylation of Lipoplex
PEGylation has been extensively used for decades in liposomology. Benefits of PEGylation are very well known and that includes improvement of systematic circulation of lipoplexes. However, the disadvantage of PEGylation of lipoplexes have been very well documented as well. This includes preventing association of the lipoplex with the target cell and the inhibition of the release of DNA from the endosomal compartments which results in very low transfection efficiency.
The cytotoxicity of the cationic liposomes increased as the molecular weight of the PEGylated lipid incorporated into the cationic liposomes increases. As an example, lipoplex containing PEG5000 was more cytotoxic than a lipoplex containing PEG2000. The transfection activity of cationic liposomes decreases with increasing the percentage of PEG-DSPE lipid. Previous studies have shown that the addition of 0.5%, 1% and 2% PEG-PE reduces the activity down to 60%, 40% and 10% of original luciferase activity in lungs, respectively. It has also been reported that adding PEG-PE (1% of cationic lipid) into the freshly formed plasmid-liposome complexes could prevent the complexes from aggregating during storage. However, storage of the complexes containing PEG-PE at 4°C slowly restored the original activity. In general, PEGylation is not used for transfection experiments. However, in some experiments especially in vivo experiment PEGylated lipoplexes are used.
If you need to do an experiment which involves PEGylation of cationic liposomes, it is highly recommended to form the lipoplex first by adding the proper amount of the genetic material into the liposomes and next add PEGylated lipid externally (post-insertion) and incubate the lipoplex and the PEG lipid for one hour at a temperature which is above the liquid to gel phase transition temperature of the cationic lipid (in case of unsaturated cationic lipids such as DOTAP, you can incubate at room temperature).
In many experiments, instead of using the traditional PEG2000-DSPE, C8-PEG2000-Ceramide is used for PEGylation of the lipoplex because PEG-Ceramide is a “shedable PEG,” which diffuse out of lipoplex upon contact with biological membranes.
Cell Viability Assay
Due to cytotoxicity of cationic lipids used in the liposome formulation, it is highly recommended to perform cell viability assay. Cell viability can be assessed by a modified Alamar blue assay. Alamar blue contains a redox indicator which exhibits both fluorescence changes in the oxidation-reduction range of cellular metabolism. Briefly, add 10 μL of 10% (v/v) Alamar blue dye to 100 μL sample. After 2 h of incubation at 37ºC, the resulting fluorescence is read on a fluorescence spectrophotometer at 570 nm and 600 nm. Cell viability (a percentage of control cells) is calculated using the following equation:
![]()
Technical Notes
- Genesome® products are formulated using deionized RNAse-free water.
- N/P ratio means nitrogen to phosphate ratio. N stands for nitrogen and NOT negative, and nitrogen is positively charged. P stands for phosphate and NOT positive. Phosphate carries negative charge. Therefore, N/P ratio also stands for positive to negative ratio. Some cationic lipids contain more than one nitrogen, but not all nitrogen atoms carry a positive charge. For example, DC-Cholesterol has two nitrogen atoms and only one of them positive charged. The nitrogen next to the carboxyl group is not positively charged and should not be included in N/P calculation.
- Cytotoxicity can also be assessed using the CytoTox 96 Non-Radioactive Cytotoxicity Assay (Promega, Madison, WI) per manufacturer’s instructions.
- Liposomes should be kept at 4°C and NEVER be frozen.
Appearance
Plain Genesome® is a white translucent liquid made of nano size unilamellar liposomes. Fluorescent Genesome® formulation is colored and the color depends on the type of the fluorescent dye that is used (see SDS for appearance). Usually due to the small size of liposomes no settling will occur in the bottom of the vial. The liposomes are packaged in an amber vial.
Ordering/Shipping Information
- All liposome based formulations are shipped on blue ice at 4°C in insulated packages using overnight shipping or international express shipping.
- Liposomes should NEVER be frozen. Ice crystals that form in the lipid membrane can rupture the membrane, change the size of the liposomes and cause the encapsulated drug to leak out. Liposomes in liquid form should always be kept in the refrigerator.
- Clients who order from outside of the United States of America are responsible for their government import taxes and customs paperwork. Encapsula NanoSciences is NOT responsible for importation fees to countries outside of the United States of America.
- We strongly encourage the clients in Japan, Korea, Taiwan and China to order via a distributor. Tough customs clearance regulations in these countries will cause delay in custom clearance of these perishable formulations if ordered directly through us. Distributors can easily clear the packages from customs. To see the list of the distributors click here.
- Clients ordering from universities and research institutes in Australia should keep in mind that the liposome formulations are made from synthetic material and the formulations do not require a “permit to import quarantine material”. Liposomes are NOT biological products.
- If you would like your institute’s FedEx or DHL account to be charged for shipping, then please provide the account number at the time of ordering.
- Encapsula NanoSciences has no control over delays due to inclement weather or customs clearance delays. You will receive a FedEx or DHL tracking number once your order is confirmed. Contact FedEx or DHL in advance and make sure that the paperwork for customs is done on time. All subsequent shipping inquiries should be directed to Federal Express or DHL.
Storage and Shelf Life
Storage
Genesome® products should always be stored at in the dark at 4°C, except when brought to room temperature for brief periods prior to animal dosing. DO NOT FREEZE. If the suspension is frozen, the encapsulated drug can be released from the liposomes thus limiting its effectiveness. In addition, the size of the liposomes will also change upon freezing and thawing.
Shelf Life
Genesome® is made on daily basis. The batch that is shipped is manufactured on the same day. It is advised to use the plain Genesome® within 4 months and fluorescent Genesome® within 2 months of the manufacturing date.











